3D replica of senecavirus, a pathogen discovered several years ago by researchers in Pennsylvania. UC San Francisco researcher Eric Delwart and his colleague Chunlin Wang of Stanford University use the RENCI-developed TeraGrid Science Gateway to access grid computing resources in their search for new viruses. (Credit: Institute for Animal Health, UK)
DNA sequencing and sequence analysis happens daily in many biological sciences laboratories, but analyzing large sets of genetic data increasingly requires computing resources beyond the capabilities of most labs.
That search for the best hardware and software led Eric Delwart, a professor of laboratory medicine at the University of California, San Francisco, and a senior investigator at the Blood Systems Research Institute, and Chunlin Wang, a research associate at the Stanford University Genome Technology Center, to the Renaissance Computing Institute’s (RENCI) Engagement Team and then to the distributed computing resources of the TeraGrid and the Open Science Grid (OSG).
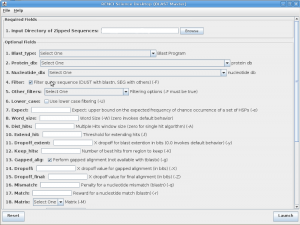
Screen shot of the BLASTMaster interface, part of the RENCI-developed TeraGrid Science Gateway interface that allows researchers to easily use distributed grid resources.
Delwart works with Wang to identify new viruses. The team uses a technique called massively parallel pyrosequencing, which can determine sequences for millions of DNA fragments using high-throughput computing. The resulting DNA sequences are then compared to all the sequences in public sequence databases to identify viral fingerprint sequences. One single sequencing reaction generates massive volumes of data that can take months, even years, to analyze on a small-scale computing cluster.
In an effort to analyze more data more efficiently, Delwart and Wang turned to the RENCI Engagement Team, which participates in the TeraGrid Science Gateways program and the leads the OSG Engagement program. TeraGrid’s Science Gateways aim to bring new communities of users to TeraGrid resources by providing easy access to the TeraGrid’s distributed computing resources. The OSG Engagement program recruits new users from a wide range of disciplines and helps them become users of the distributed computing systems operated and maintained by OSG members.
The effort to find more computing power paid off for the Delwart and Wang: In early May, they used 70,000 CPU hours on TeraGrid and OSG resources to complete in a week a DNA sequence analysis that would have taken over three months on their own lab cluster. The team submitted its jobs to the national resources using a RENCI-developed, Web services-based computational science platform [reference: http://www.teragrid.org/tg09/files/tg09_submission_75.pdf ].
“We created an application that communicates with RENCI’s TeraGrid Science Gateway,” said Jason Reilly, a RENCI senior research software developer. “For the user, it’s very simple—just log in and the application maps the data input to specific tasks to be done. The beauty is you don’t have to submit commands over and over again. You can run hundreds or even thousands of operations and you only have to submit the command once.”
The custom application created by Reilly was dubbed BLASTMaster because it builds on the Basic Local Alignment Search Tool (BLAST) used to search sequence databases. BLASTMaster divides commands into tasks and pushes the work to RENCI’s TeraGrid Science Gateway, which submits, monitors, and manages the compute workload on systems that are part of TeraGrid’s nationwide network of high performance machines, and to OSG machines. After entering the initial commands, the researchers merely had to wait for their results.
3D replica of senecavirus, a pathogen discovered several years ago by researchers in Pennsylvania. UC San Francisco researcher Eric Delwart and his colleague Chunlin Wang of Stanford University use the RENCI-developed TeraGrid Science Gateway to access grid computing resources in their search for new viruses. Credit: Institute for Animal Health, UK
“Large computer farms that we might use are often composed of heterogeneous smaller clusters,” said Wang. “The BLASTMaster tool and a Web services environment is particularly useful to those of us without much experience using compute clusters. It gives us a uniform interface to submit jobs, which greatly enhances our productiveness.”
The sequence analysis work used TeraGrid resources at Purdue University (West Layfayette, IN), OSG resources at RENCI (Chapel Hill, NC) and a cluster in the University of North Carolina at Chapel Hill computer science department supported by the National Institutes of Health. The work has real-world value taken straight from recent headlines about the H1N1 virus.
“Knowing the genomic sequence of a human virus allows for quicker diagnostics to identify infections,” said Delwart. “Quicker diagnostics can lead to more informed decisions on how an emerging virus is spread and how to control it. Knowing the sequence can also help make vaccines or anti-virals against that virus.”
_______________
OSG is a consortium of universities, national laboratories, scientific collaborations and software developers dedicated to meeting the ever-growing computing and data management requirements of scientific researchers. Supported by the U.S. Department of Energy Office of Science and the National Science Foundation, it provides access to its members’ independently owned and managed resources through a common grid infrastructure that uses high performance networks to connect computing systems scattered across the country. For more see http://www.opensciencegrid.org.
TeraGrid is an open scientific discovery infrastructure combining leadership class resources at eleven partner sites to create an integrated, persistent computational resource. TeraGrid is funded by the National Science Foundation. For more, see http://www.teragrid.org.
The Renaissance Computing Institute (RENCI), a multi-institutional organization, brings together multidisciplinary experts and advanced technological capabilities to address pressing research issues and to find solutions to complex problems that affect the quality of life in North Carolina, our nation and the world. Founded in 2004 as a major collaboration of Duke University, North Carolina State University, the University of North Carolina at Chapel Hill and the state of North Carolina, RENCI is a statewide virtual organization. For more see http://www.renci.org


